Data preprocessing and analysis
Molecular representation and description
Each molecular structure in this study was represented as a SMILES42 string. CATS56 descriptors and 2048-bit ECFPs53 (using a radius of 2) were computed for each molecule.
Data collection and curation
Thirty-three labelled datasets of molecular structures with their corresponding experimental target property were used:
Moreover, small molecules were collected from ChEMBLv3345 for model pretraining. Because molecular structures from ChEMBL were used for pretraining, molecules with a Bemis–Murcko scaffold76 similar to any such scaffold in the labelled datasets (Tanimoto similarity coefficient54 on EFCPs larger than 0.7) were removed. This included molecules without a Bemis–Murcko scaffold (that is, containing no rings).
For prospective virtual screening, the most recent CDK1 (CHEMBL308) data were fetched from ChEMBL v3545 (accessed in April 2025) as an additional and independent dataset. Raw data were processed in accordance with previous work48.
All bioactivity endpoints of the MoleculeACE48 and CDK1 datasets were converted from continuous regression labels into binary classification labels. Molecules with an EC50 (half maximal effective concentration) or Ki (inhibitory constant) of 100 nM or lower were labelled as bioactive, whereas less potent molecules were labelled as inactive. For the LIT-PCBA49 and Ames mutagenicity dataset50, their original binary classification labels were used.
For all datasets, SMILES strings were preprocessed using RDKit v. 2024.3.377. For each SMILES string, stereochemistry tokens as well as salts and solvents (Supplementary Table 2) were removed. Each molecule was sanitized, neutralized using predefined neutralization reactions (Supplementary Table 3), and its SMILES string was canonicalized. Disconnected structures and molecules that contained formal charges, contained complex ring systems (SMILES strings with a ring index of 9 or higher), non-standard isotopes or any atoms other than Cl, Br, H, C, N, O, F, S and I were removed. Molecules were removed if they contained more than 100 tokens in their canonical SMILES string or if they could not be featurized into CATS descriptors and/or ECFP fingerprints. Sizes of datasets before and after data curation are presented in Supplementary Table 1.
Virtual screening library (retrospective)
Molecules were gathered from three commercial screening libraries:
All molecules were processed in the same way as the training datasets. Finally, all unique molecules were aggregated.
Virtual screening library (prospective)
The most recent Specs library was downloaded from https://www.specs.net/index.php?view=databases&page=download (accessed in April 2025). All molecules were processed in the same way as the training datasets. To ensure that molecules were compatible with our experimental assay (likely soluble in 1% dimethylsulfoxide (DMSO)), physicochemical rules were enforced based on relaxed rule-of-five and Veber criteria:
-
(1)
Molecular weight between 200 g mol−1 and 600 g mol−1.
-
(2)
logP lower than 6.
-
(3)
Total polar surface area between 20 Å2 and 140 Å2.
-
(4)
Number of hydrogen bond donors lower than six.
-
(5)
Number of rotatable bonds lower than ten.
-
(6)
A maximum of two rule-of-five violations.
In addition, molecules with a terminal enone, isocyanate, quinone, aromatic nitro groups, azide or epoxide groups were removed by using SMILES Arbitrary Target Specification (SMARTS) patterns to prevent assay interference. Furthermore, to prevent the selection of trivially simple molecules and enrich the general-purpose Specs library for kinase-relevant chemical space, several general criteria were enforced:
-
(1)
A molecule must have an ATP mimetic core, that is, at least one heteroatom in a ring, or a fused carbocyclic system.
-
(2)
A molecule must have a polar anchor to ensure solubility or solvent interaction.
-
(3)
A molecule must have enough hydrophobic mass or planarity to fill the kinase ATP pocket (for example, gatekeeper region).
-
(4)
A molecule must have a directional H-bond donor/acceptor to ensure potential for hinge interaction.
-
(5)
A molecule must have two or more rings.
These rules are intentionally permissive for new chemotypes, and 98.6% of all kinase inhibitors that ever went into clinical trials78 pass these filters. Finally, molecules with a Tanimoto similarity on ECFPs >0.7 to any molecule in the respective target’s data (PIM1, and CDK1) were removed. This left 185,298 and 185,336 screening molecules for PIM1 and CDK1, respectively.
Molecular cyclic skeletons
Cyclic skeletons51 (core ring systems without exocyclic substituents) were extracted from molecules to serve as the most fundamental molecular scaffold representation. From each molecule, Bemis–Murcko scaffolds76 were obtained, removing peripheral substituents. Remaining double-bonded exocyclic substituents were then removed, and all atoms and bonds were made generic to obtain the final cyclic skeleton.
Data splitting
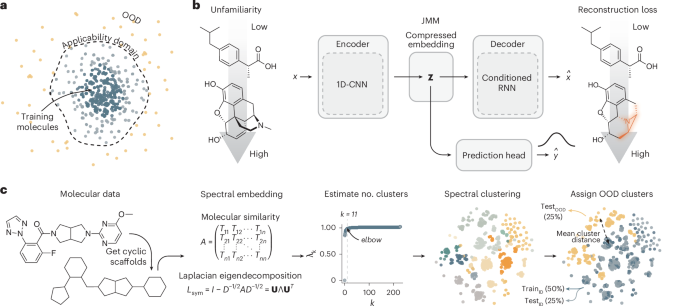
The curated molecular structures from ChEMBL were split into a training (80%, n = 1,230,041), a test set (10%, n = 153,755) and a validation (10%, n = 153,755) set, using random splitting. All labelled datasets were split into a training set (~50%), test set (~25%) and OOD set (~25%). To determine the OOD molecules, spectral clustering was performed on unique cyclic skeletons (see below). The molecules corresponding to the scaffolds in the clusters with the lowest mean cluster similarity to all other clusters that constituted approximately 25% of the total dataset size were taken as the OOD set. The remaining molecules (that is, the ~75% most similar molecules) were split randomly in a train and test set, with the test set being equal in size to the OOD set. An overview of all datasets is presented in Supplementary Table 1.
Spectral clustering
Spectral clustering was performed on a molecular similarity matrix A using Sci-kit learn79. A is an n × n matrix where each element Aij is the Tanimoto coefficient on ECFPs Tij between two molecular structures. By using a molecular similarity matrix directly, we bypass the complex, high-dimensional and non-Euclidean nature of molecular structures. From this affinity matrix, the symmetrically normalized Laplacian was constructed as follows:
$${L}_{\mathrm{sym}}=I-{D}^{-1/2}A{D}^{-1/2},$$
(3)
where I is the identity matrix and D is the degree matrix. Subsequently, eigenvalue decomposition was performed:
$${L}_{\mathrm{sym}}={\bf{U}}\Lambda {{\bf{U}}}^{T}{.}$$
(4)
To determine the number of clusters k for spectral clustering, the eigenvalues λ1, λ2, …, λn were sorted in ascending order and the elbow (the point of maximal curvature) of the resulting sequence was estimated using the kneed algorithm80. Finally, the spectral embeddings of the data were clustered by taking the top k (smallest) eigenvectors Uk, normalizing Uk to unit length and performing k-means clustering on all rows (uk).
Chemical space visualization
To visualize each labelled dataset, molecules were first encoded as ECFPs. The resulting binary ECFPs were reduced to 100 components using truncated singular value decomposition and embedded into a two-dimensional space using t-distributed stochastic neighbour embedding with a perplexity value of 30 and default settings.
MCS fraction
To compute molecular core similarity, we computed the MCS fraction55 between a molecule Ma and a reference molecule Mb as
$$\mathrm{MCS}\,\mathrm{fraction}=\frac{\left|\mathrm{MCS}\left({M}_{a},\,{M}_{b}\right)\right|}{\left|{M}_{a}\right|},$$
(5)
where |Ma| is the number of atoms in Ma and MCS is the maximal common substructure between the two molecules Ma and Mb, as determined by the FMCS algorithm7 in RDKit. A high MCS fraction indicates that a molecule shares a significant portion of their overall core structure with a reference molecule. This implementation is asymmetric.
Molecular complexity
To quantify molecular complexity we compute the well-established Bertz complexity81 and Böttcher complexity82. In addition, to align complexity measures with the task of reconstructing SMILES strings, we also estimated the complexity of the molecular graph directly83 as
$${C}_{\mathrm{graph}}=V{\log }_{2}V-\mathop{\sum }\limits_{i\in {\mathscr{G}}}{V}_{i}{\log }_{2}{p}_{i},$$
(6)
where G represents the molecular graph, V is the total number of elements in the graph and Vi is the number of the elements in the ith set of elements. In a similar fashion, we also estimated the complexity of a SMILES string, via their entropy computed as
$${C}_{\mathrm{SMILES}}=-\mathop{\sum }\limits_{i\in S}{p}_{i}{\log }_{2}{p}_{i},$$
(7)
where S represents the set of unique tokens in a SMILES string, and pi is the probability of the ith token occurring in S. Tokens representing the start, end of sequence, and padding were not considered. Moreover, for each molecule, we counted the number of SMILES tokens (excluding padding), the number of SMILES string branches (that is, ‘(‘ tokens), and the presence of 50 unique molecular patterns74.
Virtual screening
Top-k molecules were selected in a multi-objective manner by selecting the k molecules closest to the utopia point68. The distance to this ideal point can be calculated as
$${d}_{\mathrm{utopia}}=\sqrt{\mathop{\sum }\limits_{i=1}^{n}{\left({\mathrm{norm}}_{i}\right)}^{2}},$$
(8)
where normi is the normalized objective that is either maximized (for example, predicted bioactivity) or minimized (for example, prediction uncertainty):
$$\genfrac{}{}{0ex}{}{{{\rm{norm}}}_{i}}{[\text{max}]}=\frac{{x}_{\text{max}}-{x}_{i}}{{x}_{\text{max}}-{x}_{\text{min}}}{\rm{or}}\genfrac{}{}{0ex}{}{{{\rm{norm}}}_{i}}{[\text{min}]}=\frac{{x}_{i}-{x}_{\text{min}}}{{x}_{\text{max}}-{x}_{\text{min}}}.$$
(9)
Machine learning
Encoder
Canonical SMILES strings were encoded by a one-dimensional convolutional neural network. SMILES string character tokens were embedded using a randomly initialized trainable embedding layer of size 128. Several one-dimensional convolutional layers were used with a stride of 1 and no padding. Each layer was followed by a ReLU activation, standard max pooling with a kernel size equal to that of the convolutional layers, and dropout. Both convolutional and pooling layers used a stride of 1 and no padding. The final output was flattened and compressed to a latent vector (z) of size 128 using a fully connected layer. The following hyperparameters were optimized (see ‘Hyperparameter optimization’ section): the number of convolutional layers [2, 3], filter size [256, 512], kernel size [6, 8], weight decay on convolutional neural network weights [0, 1 × 10−4] and dropout [0, 0.1].
Decoder
Encoded latent molecular representations in z were reconstructed back to SMILES strings using a conditioned LSTM model. A randomly initialized trainable embedding layer of 128 neurons was used to embed SMILES string character tokens. The following hyperparameters were optimized (see ‘Hyperparameter optimization’ section): the number of LSTM layers (nlayers) [2, 3] and the LSTM hidden size (\({\mathrm{size}}_{\mathrm{layers}}\)) [256, 512]. Models were trained autoregressively without teacher forcing using next token prediction based on the tokens predicted in the previous steps rather than the ground truth tokens. A reconstruction loss normalized for sequence length was used (equation (1)). To condition the model, the LSTM hidden state h0 was initialized with z for every molecule. To correctly match all dimensions of h0 (\({n}_{\mathrm{layers}},{\mathrm{size}}_{\mathrm{layers}}\)), z was first transformed to \({n}_{\mathrm{layers}}\,\times {\mathrm{size}}_{\mathrm{layers}}\) with a fully connected layer, after which it was reshaped into \({{\rm{n}}}_{\mathrm{layers}}\) chunks of \({\mathrm{size}}_{\mathrm{layers}}\).
Approximate Bayesian classifier
Labels were predicted from either ECFPs or SMILES strings encoded into latent molecular representationsz. A MLP was used with several fully connected layers and an output layer consisting of two neurons. The number of MLP layers [2, 3] and the MLP hidden size [1,024, 2,048] were optimized (see ‘Hyperparameter optimization’ section). To estimate prediction uncertainty, anchored ensembling47 was implemented on the MLP as in our previous work74. We used an ensemble of M = 10 models. For each model, \(m\in [1\ldots M]\), we regularized its parameters θm with a set of ‘anchored’ parameters θanchor,m that prevent different models in the ensemble to converge to the same parameter space. Each model is initiated with distinct θanchor, which is controlled by different random seeds. The classification loss in our implementation is defined as
$${{\mathscr{L}}}_{\mathrm{MLP}}=-\frac{1}{M}\mathop{\sum }\limits_{m=1}^{M}\mathop{\underbrace{\log {p}_{m}\left(\,{y|x}\right)s}}\limits_{\text{prediction}}+\mathop{\underbrace{\lambda {{||}{\theta }^{m}-{\theta }_{\mathrm{anchor}}^{m}{||}}^{2}}}\limits_{\mathrm{anchoring}},\,$$
(10)
where λ is a regularization coefficient (set to 3 × 10−4). To estimate the expected value \({\mathbb{E}}\) of each molecule x, we take the mean prediction over the whole ensemble, as follows:
$${\mathbb{E}}\left(\,{y|x}\right)=\frac{1}{M}\mathop{\sum }\limits_{m=1}^{M}{p}_{m}\left(\,y,|,x\right).$$
(11)
Similarly, we estimate prediction uncertainty for each molecule \(x\) as the mean entropy \({\mathbb{H}}\) over all models in the ensemble:
$${{{\mathbb{H}}}}\left({y|x}\right)=-\frac{1}{M}\mathop{\sum }\limits_{m=1}^{M}{p}_{m}\left(\,{y|x}\right)\log {p}_{m}\left(\,{y|x}\right).$$
(12)
JMM
Canonical SMILES strings were encoded into compressed latent molecular representations \({\bf{z}}\) using the encoder model described above. Subsequently, \({\bf{z}}\) was used to perform both molecular property prediction with a classifier and molecular reconstruction using the \({\bf{z}}\)-conditioned decoder model27. The model was optimized in a joint fashion using the following weighted composite loss function:
$${{\mathscr{L}}}_{\mathrm{JMM}}={{\mathscr{L}}}_{\mathrm{reconstruction}}+\gamma \times {{\mathscr{L}}}_{\mathrm{MLP}},$$
(13)
where the scalar γ was set at 0.1 based on preliminary experiments. A regular autoencoder was used as preliminary experiments showed no performance benefits of the more complex variational autoencoder84 architecture.
RF
An RF classifier was trained on either ECFPs or CATS56 descriptors. The following hyperparameters were optimized (see ‘Hyperparameter optimization’): the number of trees [100, 250, 500, 1,000], the maximal tree depth [10, 20, 30, ∞] and the minimal samples per split [2, 5, 10].
Model training
Autoencoders
Encoder–decoder models were (pre)trained to reconstruct SMILES strings of general drug-like molecules from ChEMBL using the Adam optimizer. Mini batches of 256 random molecules were sampled from the training data for 1,000,000 steps using uniform sampling. Gradients were clipped with a max norm of 5. Early stopping with a patience of up to 20 evaluation checkpoints was implemented by monitoring validation loss every 10,000 steps. The model checkpoint with the best validation loss was used.
Classifiers
Classifiers using ECFPs or SMILES strings as input were trained for molecular property prediction on each of the labelled dataset using a similar setup to the autoencoders. However, tenfold Monte Carlo cross-validation was used with 10% validation splits instead of one predefined data split. Mini batches of 64 were resampled during training based on the occurrence of their class with
$${P}_{c}=1-\frac{{n}_{c}}{N},$$
(14)
where Pc is the probability of sampling class c, nc is the number of samples of class c, and N is the total number of samples. Models were trained for 5,000 steps with an early stopping patience of 10 evaluation checkpoints, performed every 10 steps. The model checkpoint with the best validation loss was used. For the RF control models, molecules were weighted inversely proportionally to their class frequency to mitigate class imbalance during training.
JMMs
Joint models, each consisting of a SMILES string encoder, a decoder and a classifier (Fig. 1b), were initialized with pretrained weights. The SMILES string encoders and classifiers used weights from models trained on the labelled datasets. For the decoder, decoder weights were used from an autoencoder pretrained on ChEMBL. Using a mini batch size of 64, the joint models were finetuned for 10,000 steps with an early stopping patience of 50 evaluation checkpoints, performed every 20 steps. A learning rate of 3 × 10−6 was used for the encoder and classifier, whereas a learning rate of 3 × 10−7 was used for the decoder. No weight decay was applied.
Hyperparameter optimization
Hyperparameters, as specified previously, were optimized for all autoencoders, classifiers (using ECFPs or SMILES strings), and RF control models using a simple grid search. Tenfold Monte Carlo cross-validation was used, repeatedly using 10% of the training data as a validation split. The hyperparameters of the model with the best mean validation loss was used.
Model evaluation
Predictions were evaluated according to the following performance metrics:
$$\text{Balanced accuracy}=\frac{1}{2}\left(\frac{\mathrm{TP}}{\mathrm{TP}+\mathrm{FN}}+\frac{\mathrm{TN}}{\mathrm{TN}+\mathrm{FP}}\right),$$
(15)
$$\text{Precision}=\frac{\mathrm{TP}}{\mathrm{TP}+\mathrm{FP}},$$
(16)
where TP is the number of true (that is, correctly predicted) positives, TN is the number of true negatives, FN is the number of false negatives, and FP is the number of false positives. In addition, the hit rate (true positive rate) was determined for virtual screening experiments as
$$\text{Hit rate}=\frac{\mathrm{TP}}{P},$$
(17)
and enrichment factor as
$$\text{Enrichment factor}=\frac{{\mathrm{TP}}_{k}}{P/N}.$$
(18)
Here, TPk is the number of correctly identified positives in the subset of k prioritized molecules, P is the total number of positives in the full dataset, and N is the total number of molecules in the full dataset.
Biological characterization
Sixty screening compounds were purchased from Specs Compound Handling B.V. and dissolved at 10 mM in 100% DMSO.
To screen for bioactivity, a point screening was first performed at a concentration of 10 µM (in 1% DMSO) in technical triplicates using the ADP-Glo Kinase Assay platform from Promega using the Chemi-Verse PIM1 Kinase Assay Kit and the Chemi-Verse CDK1/CyclinA2 Kinase Assay Kit from BPS Bioscience in Costar flat white 96-well plates. AZD1208 (CAS 1204144-28-4) and dinaciclib (CAS 779353-01-4), purchased from TargetMol Chemicals, were used as positive controls for PIM1 and CDK1, respectively. Bioactivity was measured as the area under the curve of an 18-step luminescence scan between 398 nm and 653 nm with an integration time of 1 s and a settle time of 100 ms, normalized for the signal in buffer-only wells.
For each target protein, the six compounds with the highest bioactivity at 10 µM were followed up with an 8-point dose–response curve using the same assay. Screening compounds were measured in technical triplicates from 10 µM to 0.0046 µM, whereas reference compounds were measured in duplicate.
Hardware and training set-up
All computational experiments were performed on a Lenovo ThinkSystem SD650-N v2 server equipped with Intel Xeon Platinum 8360Y central processing units and NVIDIA A100 (40 GB) graphics processing units. Up to five models were trained in parallel on a single graphics processing unit.
Software and code
All code was implemented in Python (v. 3.12). Deep learning models were implemented using PyTorch (v. 2.3.0)85. Traditional machine learning models and clustering was implemented using Sci-kit learn v.1.4.0 (ref. 79). All molecular data were handled using RDKit (v.2024.3.3)77. For data visualization, R (v.4.3.0) and the R package ggplot2 (v.3.4.2) were used along with Adobe Illustrator.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.